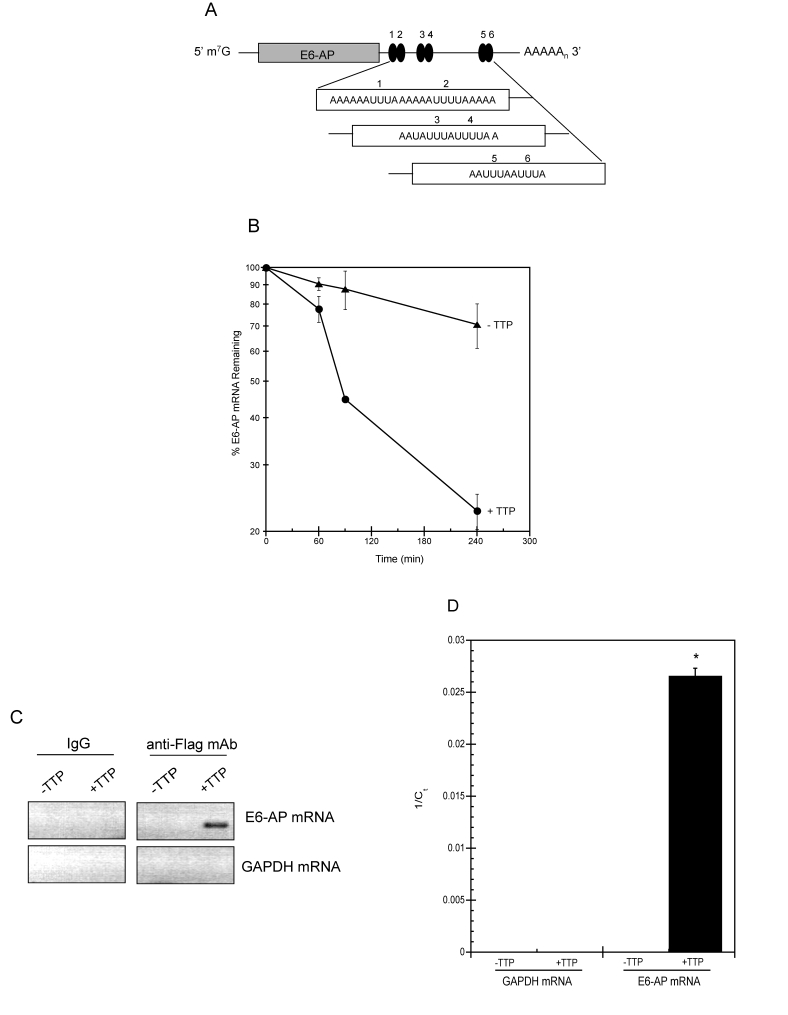
Figure 6.TTP binds E6-AP mRNA and targets it for rapid decay.
(A) Schematic representation of E6-AP mRNA. The grey bar
corresponds to E6-AP coding region and number-labeled black ovals
represent putative 3' UTR AU-rich elements (AREs). m7G, 7-methyl-guanosine
cap; AAAAn, polyadenylated tail. (B) E6-AP mRNA half-life
was assayed in HeLa Tet-Off/TTP-Flag cells grown in the presence
(triangles; labeled -TTP) or absence (circles; labeled +TTP) of
Dox to induce TTP expression. After 48 hr, 5 μg/ml of actinomycin
D was added to the cells and E6-AP mRNA decay was analyzed by qPCR
using GAPDH mRNA as a normalization control. The data shown is the
average of triplicate experiments. (C, D) Binding of TTP
and E6-AP mRNA. Control and TTP-expressing (48 hr) HeLa Tet-Off/TTP-Flag
cells were lysed and immunoprecipitation was performed on equal
amounts of cytoplasmic lysates using control IgG or anti-Flag mAb.
RNA purified from immuno-precipitates was subjected to RT-PCR (C)
or qPCR (D) to detect E6-AP and GAPDH mRNA. The ethidium
bromide-stained agarose gel depicting the 292bp E6-AP PCR product
is shown in reverse image. The relative amounts of immuno-precipitated
E6-AP mRNA is reported as the average 1/Ct value of triplicate experiments.
(*) P < 0.01